|
I am a research scientist at Genentech, Prescient Design team. Previously, I was a research scientist at Deep Genomics and Element AI (where I work closely with people from Mila). I received a Ph.D. from École Polytechnique Fédérale de Lausanne (EPFL) and Idiap Research Institute (Switzerland), under supervision of Ronan Collobert. During my phd, I also spent time at Facebook AI Research (FAIR). Previously, I graduated with a MSc. in Image and Signal Processing from Institut National des Sciences Appliquées de Lyon (INSA), in France. I am originally from sunny Fortaleza, Brazil. |

|
|
I am interested in machine learning and its applications, specifically deep learning methods for representation learning and generative modeling.
Currently, I am interested in developing AI methods that can tackle complex scientific problems, such as molecular sciences and drug discovery.
Previously, my work focused on computer vision applications.
|
|
Pedro O. Pinheiro, Pan Kessel, Aya A. Ismail, Sai P. Mahajan, Kyunghyun Cho, Saeed Saremi, Nataša Tagasovska NeurIPS, 2025 paper |

|
Matthieu Kirchmeyer*, Pedro O. Pinheiro*, Karolis Martinkus, Emma Willett, ..., Richard Bonneau, Saeed Saremi NeurIPS, 2025 (*equal contribution) paper | code |
|
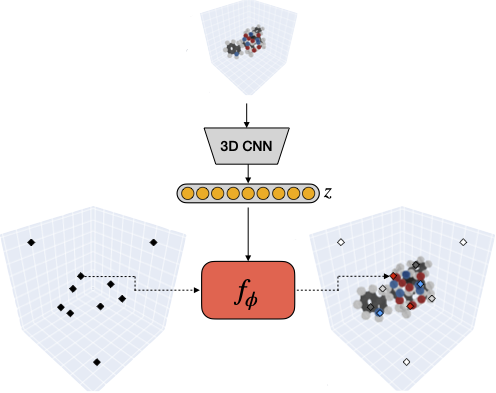
Matthieu Kirchmeyer*, Pedro O. Pinheiro*, Saeed Saremi NeurIPS, 2024 (*equal contribution) paper | code |
|
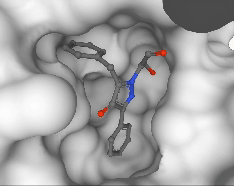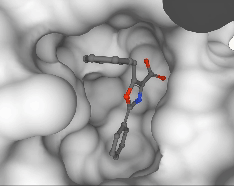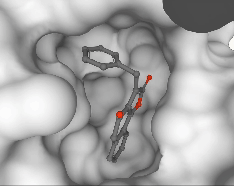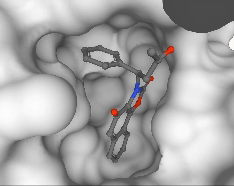
Pedro O. Pinheiro, Arian Jamasb, Omar Mahmood, Vishnu Sresht, Saeed Saremi ICML, 2024 paper | code |
|
Pedro O. Pinheiro, Joshua Rackers, Joseph Kleinhenz, Michael Maser, Omar Mahmood, Andrew Martin Watkins, Stephen Ra, Vishnu Sresht, Saeed Saremi NeurIPS, 2023 paper | code |
|
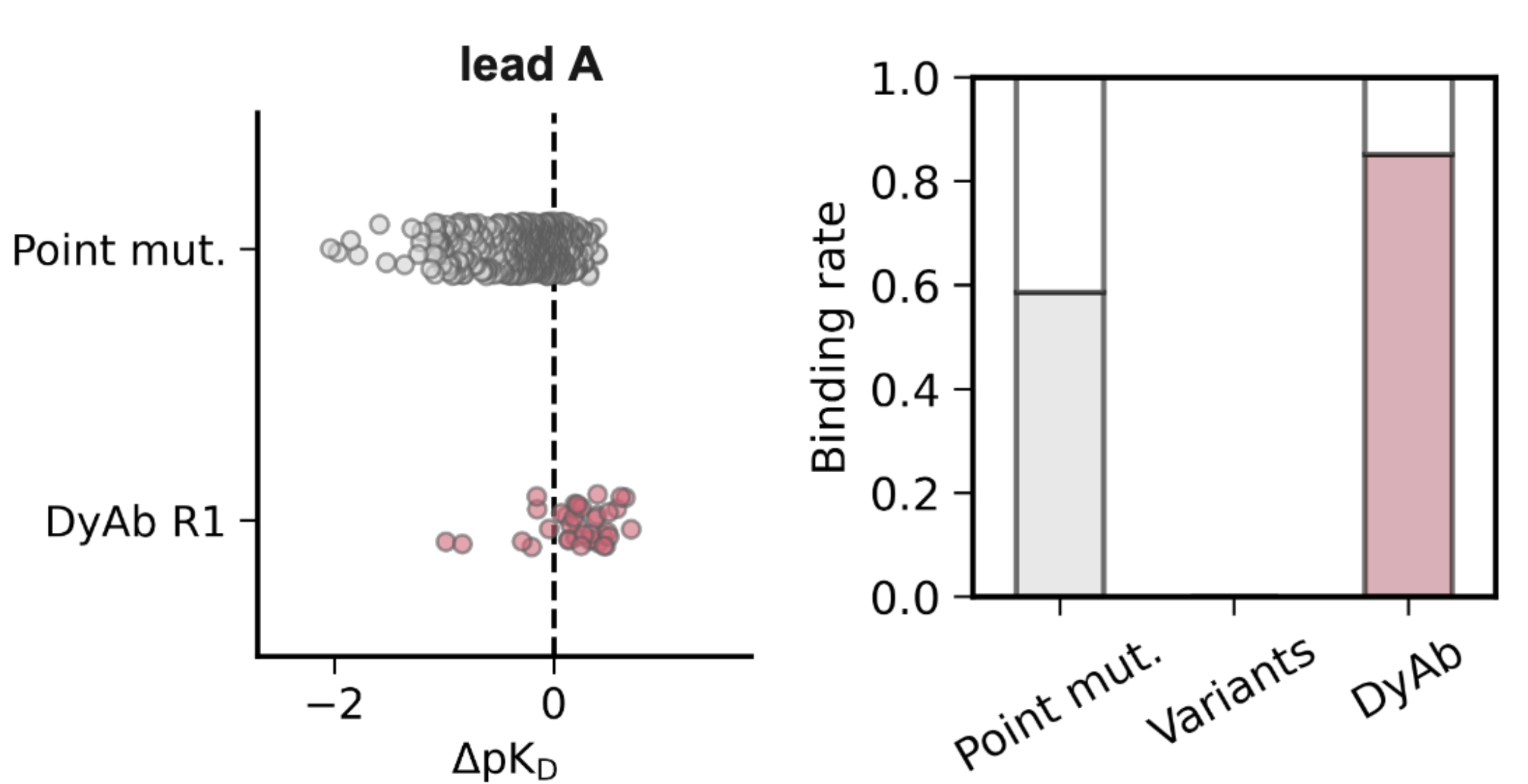
Joshua Yao-Yu Lin, Jennifer L. Hofmann, Andrew Leaver-Fay,..., Pedro O. Pinheiro,..., Andrew Watkins, Kyunghyun Cho, and Nathan C. Frey bioRxiv, 2025 paper |

|
Albi Celaj, Alice Jiexin Gao, Tammy T.Y. Lau, Erle M. Holgersen, ..., Pedro O. Pinheiro, ..., Brendan J. Frey bioRxiv, 2023 paper |

|
Pedro O. Pinheiro, Amjad Almahairi, Ryan Benmalek, Florian Golemo, Aaron Courville NeurIPS, 2020 paper |

|
Sai Rajeswar, Cyril Ibrahim, Nitin Surya, Florian Golemo, David Vazquez, Aaron Courville, Pedro O. Pinheiro CoRL , 2021 paper | code |

|
Arantxa Casanova, Pedro O. Pinheiro, Negar Rostamzadeh, Christopher J. Pal ICLR, 2020 paper | code |

|
Scott Weichenthal, Evi Dons, Kris Hong, Pedro O. Pinheiro, Filip JR Meysman Environmental Research, 2020 paper |

|
Kris Hong, Pedro O. Pinheiro, Scott Weichenthal Environmental International, 2020 paper |

|
C Xing, N Rostamzadeh, B Oreshkin, Pedro O. Pinheiro NeurIPS, 2019 paper | code |

|
Jae Hyun Lim, Pedro O. Pinheiro, Negar Rostamzadeh, Christopher Pal, Sungjin Ahn Neurips , 2019 paper | code |

|
Pedro O. Pinheiro, Negar Rostamzadeh, Sungjin Ahn ICCV, 2019 paper |

|
Kris Hong, Pedro O. Pinheiro, Laura Minet, Marianne Hatzopoulou, Scott Weichenthal Environmental Research, 2019 paper |

|
Pedro O. Pinheiro CVPR, 2018 paper |

|
Issam Laradji, Negar Rostamzadeh, Pedro O. Pinheiro, David Vazquez, Mark Schmidt ECCV, 2018 paper | code |

|
Pedro O. Pinheiro, Tsung-Yi Lin, Ronan Collobert, Piotr Dollár ECCV, 2016 paper | code |

|
Pedro O. Pinheiro, Ronan Collobert, Piotr Dollár NIPS, 2015 paper | code |

|
Pedro O. Pinheiro, Ronan Collobert CVPR, 2015 paper |

|
Rémi Lebret*, Pedro O. Pinheiro*, Ronan Collobert ICML, 2015 paper |

|
Pedro O. Pinheiro, Ronan Collobert ICML, 2014 paper |